Probe CUST_12282_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12282_PI426222305 | JHI_St_60k_v1 | DMT400043145 | GTCTCGAGCAATCGGTAAGCATATTAGTAGTTAAATGGTCAATGAGGATAATTTAACTAG |
All Microarray Probes Designed to Gene DMG400016742
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12282_PI426222305 | JHI_St_60k_v1 | DMT400043145 | GTCTCGAGCAATCGGTAAGCATATTAGTAGTTAAATGGTCAATGAGGATAATTTAACTAG |
| CUST_12218_PI426222305 | JHI_St_60k_v1 | DMT400043146 | TAATCTATTGGGATGGTGCAAGAGTTCTTGGAGTTTTGGCTATGTCTCGAGCAATCGGTG |
Microarray Signals from CUST_12282_PI426222305
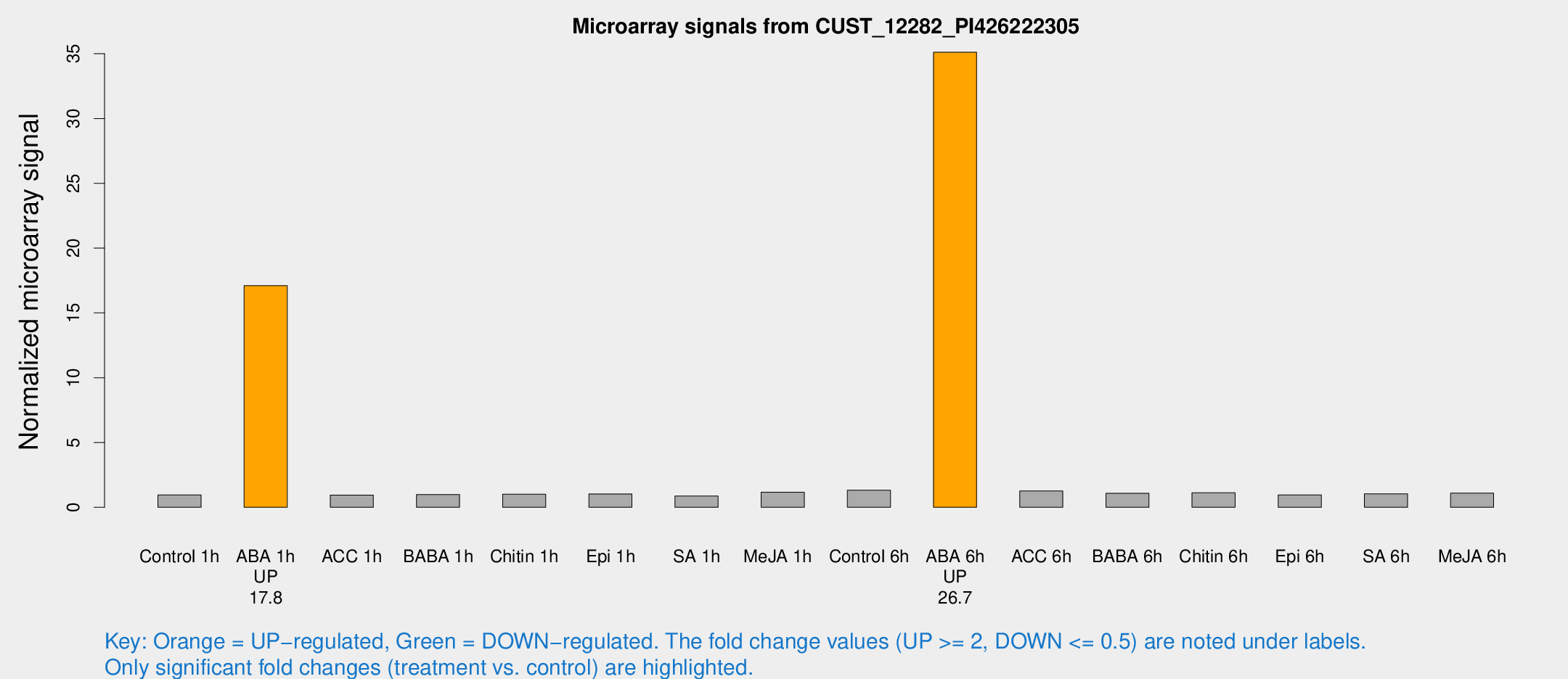
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.49599 | 3.43221 | 0.960965 | 0.514782 |
| ABA 1h | 112.123 | 34.9497 | 17.1014 | 5.37099 |
| ACC 1h | 6.55007 | 3.80217 | 0.947277 | 0.548583 |
| BABA 1h | 6.40561 | 3.71022 | 0.989053 | 0.572717 |
| Chitin 1h | 6.10559 | 3.5488 | 1.00789 | 0.584322 |
| Epi 1h | 5.99841 | 3.48366 | 1.02996 | 0.597019 |
| SA 1h | 5.97777 | 3.46384 | 0.865738 | 0.501335 |
| Me-JA 1h | 6.43738 | 3.57301 | 1.17237 | 0.65031 |
| Control 6h | 9.12755 | 3.99706 | 1.31508 | 0.602203 |
| ABA 6h | 271.508 | 67.021 | 35.1149 | 9.32253 |
| ACC 6h | 10.2096 | 4.53395 | 1.25792 | 0.604697 |
| BABA 6h | 8.18474 | 4.14571 | 1.07814 | 0.557961 |
| Chitin 6h | 8.18193 | 4.03867 | 1.12175 | 0.57709 |
| Epi 6h | 7.3053 | 4.26678 | 0.956401 | 0.553878 |
| SA 6h | 6.87518 | 3.88149 | 1.03378 | 0.584276 |
| Me-JA 6h | 7.51616 | 3.71927 | 1.09382 | 0.565881 |
Source Transcript PGSC0003DMT400043145 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G11410.1 | +3 | 2e-93 | 296 | 179/375 (48%) | protein phosphatase 2CA | chr3:3584181-3585649 REVERSE LENGTH=399 |
