Probe CUST_12208_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12208_PI426222305 | JHI_St_60k_v1 | DMT400010462 | CAAATATCACTGCTTTCTCTGGGAGGCTTCTCAAGACAGCTTTCATTGAGGTAAACACAA |
All Microarray Probes Designed to Gene DMG400004087
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_12208_PI426222305 | JHI_St_60k_v1 | DMT400010462 | CAAATATCACTGCTTTCTCTGGGAGGCTTCTCAAGACAGCTTTCATTGAGGTAAACACAA |
| CUST_12170_PI426222305 | JHI_St_60k_v1 | DMT400010463 | GTGGTGTTTTTCACAAGAAATCTAAGTCAATGAAAGTCTACCTTCATGTCCTTTTCTAAC |
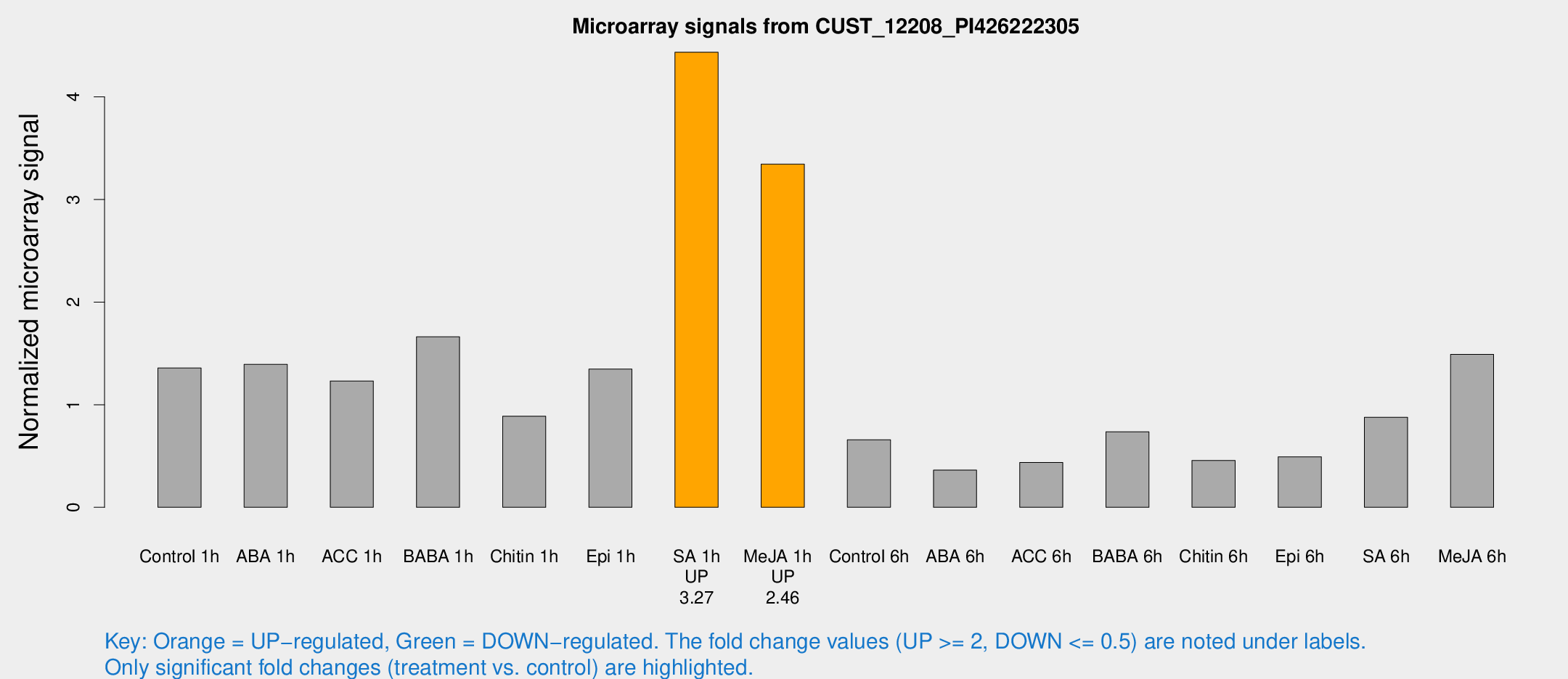
Microarray Signals from CUST_12208_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6098.56 | 671.099 | 1.35711 | 0.14766 |
| ABA 1h | 5629.69 | 970.766 | 1.39286 | 0.140876 |
| ACC 1h | 6211.06 | 1877.22 | 1.23152 | 0.340455 |
| BABA 1h | 7200.26 | 792.856 | 1.66146 | 0.0959285 |
| Chitin 1h | 3579.04 | 367.04 | 0.888081 | 0.05128 |
| Epi 1h | 5194.45 | 350.501 | 1.34656 | 0.104531 |
| SA 1h | 21402.4 | 5329.48 | 4.43499 | 1.05439 |
| Me-JA 1h | 12311.8 | 1610.22 | 3.34425 | 0.260474 |
| Control 6h | 3355.17 | 1123.69 | 0.658294 | 0.20632 |
| ABA 6h | 1721.82 | 148.529 | 0.362279 | 0.0209306 |
| ACC 6h | 2250.95 | 260.208 | 0.436347 | 0.0883239 |
| BABA 6h | 3678.02 | 288.97 | 0.735864 | 0.0816136 |
| Chitin 6h | 2162.62 | 135.778 | 0.456091 | 0.0328977 |
| Epi 6h | 2472.47 | 176.579 | 0.491721 | 0.0545218 |
| SA 6h | 3964.5 | 627.333 | 0.877508 | 0.0506707 |
| Me-JA 6h | 6760.84 | 1057.8 | 1.49012 | 0.334925 |
Source Transcript PGSC0003DMT400010462 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G29930.3 | +2 | 5e-31 | 119 | 75/155 (48%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr4:14644108-14645505 FORWARD LENGTH=263 |
