Probe CUST_11308_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11308_PI426222305 | JHI_St_60k_v1 | DMT400078542 | ACGAGAAGCTACTGATATATGACTTCATTCCAAATGGAAACCTTACCACCGCAATTCACG |
All Microarray Probes Designed to Gene DMG400030572
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_11038_PI426222305 | JHI_St_60k_v1 | DMT400078541 | ACGAGAAGCTACTGATATATGACTTCATTCCAAATGGAAACCTTACCACCGCAATTCACG |
| CUST_11308_PI426222305 | JHI_St_60k_v1 | DMT400078542 | ACGAGAAGCTACTGATATATGACTTCATTCCAAATGGAAACCTTACCACCGCAATTCACG |
Microarray Signals from CUST_11308_PI426222305
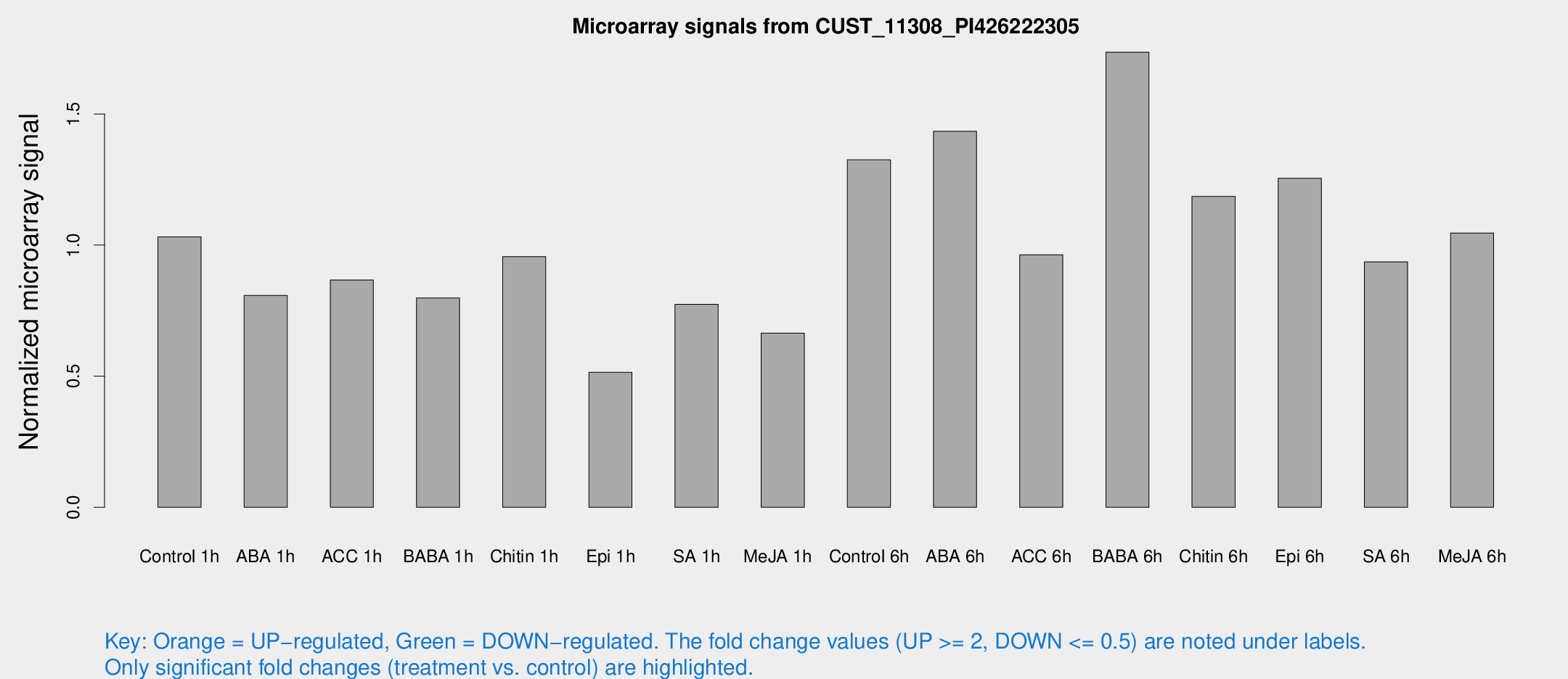
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 130.37 | 31.4927 | 1.03113 | 0.173486 |
| ABA 1h | 89.694 | 19.8015 | 0.807806 | 0.137735 |
| ACC 1h | 123.098 | 39.2236 | 0.866598 | 0.310528 |
| BABA 1h | 95.066 | 17.198 | 0.798183 | 0.108625 |
| Chitin 1h | 103.484 | 7.05825 | 0.955874 | 0.0838891 |
| Epi 1h | 62.6092 | 22.3456 | 0.514993 | 0.236548 |
| SA 1h | 97.5684 | 14.3531 | 0.773895 | 0.0747076 |
| Me-JA 1h | 65.2135 | 5.41196 | 0.664409 | 0.0923492 |
| Control 6h | 198.469 | 73.1819 | 1.32515 | 0.662742 |
| ABA 6h | 184.632 | 18.2794 | 1.43365 | 0.18287 |
| ACC 6h | 148.556 | 48.5524 | 0.963317 | 0.224096 |
| BABA 6h | 235.897 | 27.3204 | 1.73553 | 0.260965 |
| Chitin 6h | 151.545 | 9.78077 | 1.18537 | 0.0765105 |
| Epi 6h | 179.602 | 44.8456 | 1.25486 | 0.327252 |
| SA 6h | 111.47 | 7.64543 | 0.935681 | 0.0726535 |
| Me-JA 6h | 136.897 | 36.5031 | 1.04619 | 0.261273 |
Source Transcript PGSC0003DMT400078542 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G01210.1 | +3 | 0.0 | 856 | 458/703 (65%) | Leucine-rich repeat protein kinase family protein | chr2:119509-121734 REVERSE LENGTH=716 |
