Probe CUST_10367_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10367_PI426222305 | JHI_St_60k_v1 | DMT400008245 | CACATGCAAATTTGAATCTTGAATGTACGTGGAGCTGAGGATTATAGTATTGACTTGTAA |
All Microarray Probes Designed to Gene DMG400003181
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_10367_PI426222305 | JHI_St_60k_v1 | DMT400008245 | CACATGCAAATTTGAATCTTGAATGTACGTGGAGCTGAGGATTATAGTATTGACTTGTAA |
Microarray Signals from CUST_10367_PI426222305
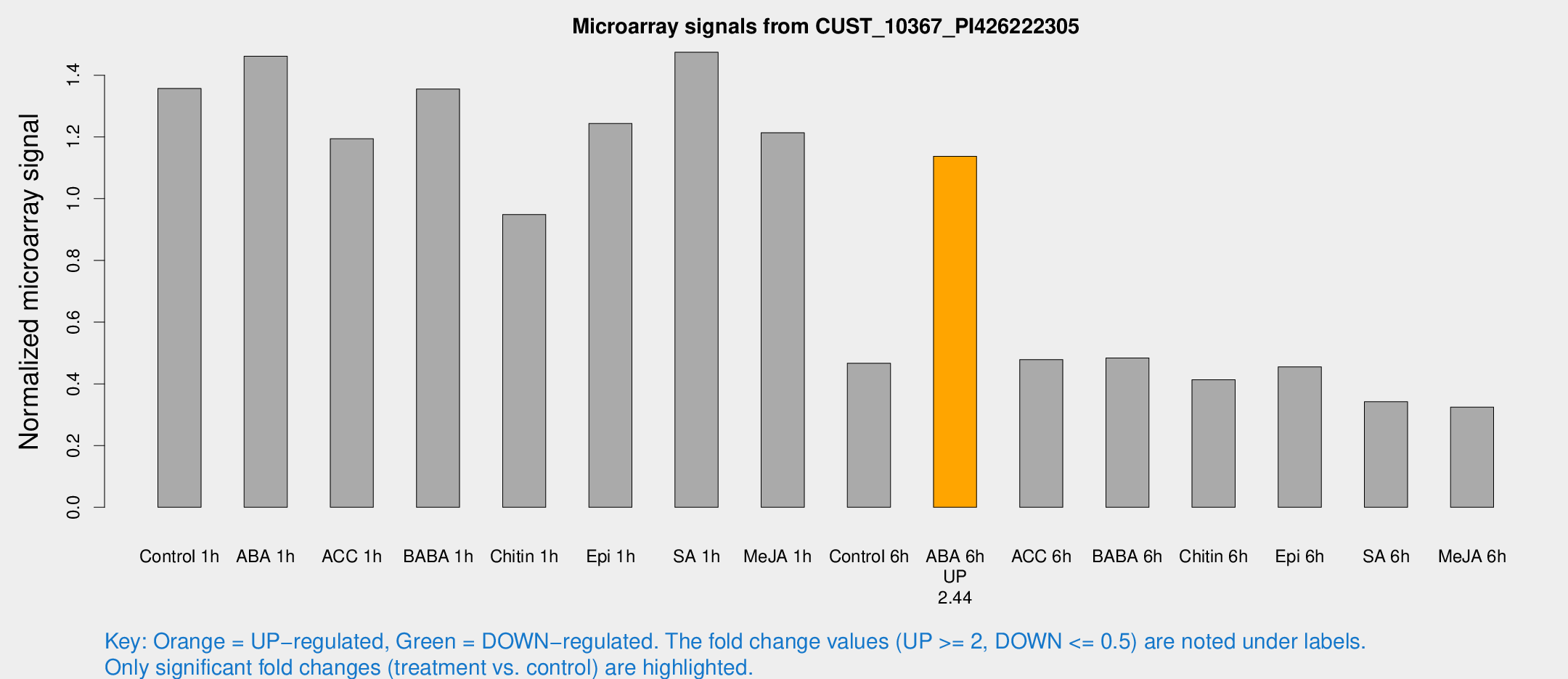
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 10803.6 | 1816.7 | 1.35691 | 0.139269 |
| ABA 1h | 10129.7 | 998.703 | 1.46117 | 0.0843621 |
| ACC 1h | 10353.9 | 2626.66 | 1.19429 | 0.287943 |
| BABA 1h | 10221.5 | 960.048 | 1.35512 | 0.0782394 |
| Chitin 1h | 6716.72 | 787.056 | 0.948597 | 0.0595792 |
| Epi 1h | 8405.68 | 672.421 | 1.24371 | 0.0788237 |
| SA 1h | 12225.8 | 2354.4 | 1.47433 | 0.257298 |
| Me-JA 1h | 7708.05 | 446.467 | 1.21333 | 0.0700536 |
| Control 6h | 3949.54 | 1032.44 | 0.466969 | 0.100843 |
| ABA 6h | 9472.27 | 971.523 | 1.13727 | 0.0656614 |
| ACC 6h | 4280.13 | 309.617 | 0.478599 | 0.0334086 |
| BABA 6h | 4231.27 | 361.099 | 0.483895 | 0.0340827 |
| Chitin 6h | 3470.81 | 452.58 | 0.413491 | 0.0507984 |
| Epi 6h | 4013.88 | 371.757 | 0.455216 | 0.0530783 |
| SA 6h | 2816.74 | 689.802 | 0.342055 | 0.0579756 |
| Me-JA 6h | 2599.73 | 511.887 | 0.324774 | 0.0444715 |
Source Transcript PGSC0003DMT400008245 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G55430.1 | +2 | 1e-152 | 448 | 232/446 (52%) | O-Glycosyl hydrolases family 17 protein | chr3:20549806-20552004 REVERSE LENGTH=449 |
